Megachile rotundata: Difference between revisions
Jump to navigation
Jump to search
| Line 484: | Line 484: | ||
s_3 35,548,153 15,521,020 4,589,276 37,564 651,924 | s_3 35,548,153 15,521,020 4,589,276 37,564 651,924 | ||
s_2_1kb 1,466,112 8,014,900 84,970 191846 | |||
lib mates mated% single% single.diffScaff% single.sameScaff% | lib mates mated% single% single.diffScaff% single.sameScaff% | ||
s_2_3kb 21,563,283 3.58 19.59 0.79 2.79 | s_2_3kb 21,563,283 3.58 19.59 0.79 2.79 | ||
Revision as of 04:30, 23 September 2010
Data
Original Traces
- 8 pairs of data files (paired ends)
cat trace.count | grep _1_ | sed 's/_sequence.txt//' | perl -ane 'print " ",$F[1],"\t",$F[0]/4,"\t",$F[0]/2,"\n";'
lib insert mates reads readLen ~coverage(500M genome) s_2_3kbp 3000 21,563,283 43,126,566 124 11 s_2_5kbp 5000 36,218,589 72,437,178 35 5 s_2_8kbp 8000 198377 396,754 124 0.1 s_3 475 35548153 71,096,306 124 18 s_4 475 35471044 70,942,088 124 18 s_5 475 35616846 71,233,692 124 18 s_6 475 35303840 70,607,680 124 18 s_7 475 34893313 69,786,626 124 18 total . 198,594,856 397,189,712 128 98*
- 2 new libs (adaptor free)
s_2_1kb 1100(10%std) 32,634,858 ? 1500 coming at the end of Sept
Corrected Traces
- Mated ones
lib insert mates reads repeatReads s_2_3kb 3000 4,823,235 9,646,470 4,349,208 (45%) s_2_8kb 8000 111,267 222,534 167,246 (75%) s_3 475 33,024,597 66,049,194 35,777,342 (54%) s_4 475 33,237,593 66,475,186 s_5 475 33,150,790 66,301,580 s_6 475 33,223,371 66,446,742 s_7 475 32,647,890 65,295,780 total . 170,218,743 340,437,486
- repeatReads:
- at least one of the mate contains a perfect match of one of the 15 frequent 22mers listed below
- 32.5%GC in repeatREads vs ~ 35.5%GC in uniqueReads
Adaptors
>circularizarion CGTAATAACTTCGTATAGCATACATTATACGAAGTTATACGA >circularizarion.revcomp TCGTATAACTTCGTATAATGTATGCTATACGAAGTTATTACG
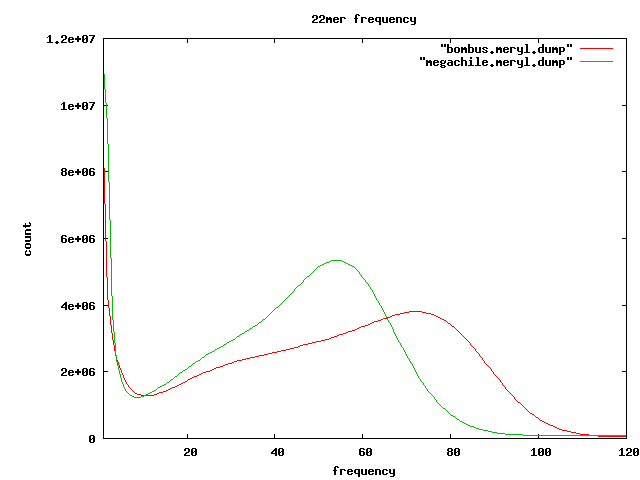
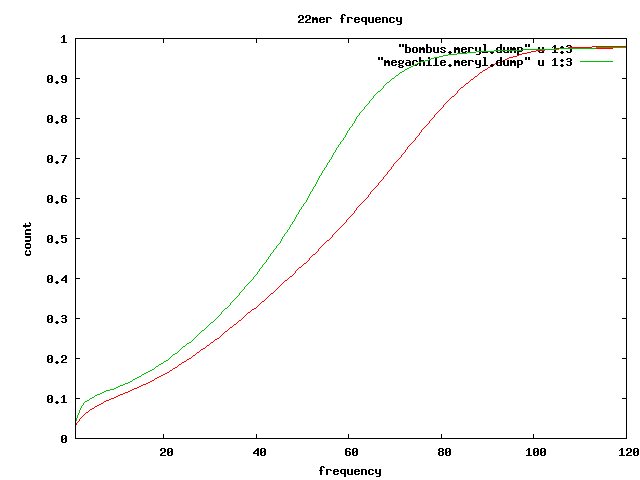
Frequent kmers
- 22mers which seem to appear in tandem
~ %reads
s_2_3kb s_2_8kb s_[4567]
-------------------------
1 AATCATACAATCACAATCATAC|GTATGATTGTGATTGTATGATT 12.04 20.1 9.99 # 14mer tandem repeat : AATCATACAATCAC|GTGATTGTATGATT
2 CAATCACAATCATACAATCACA|TGTGATTGTATGATTGTGATTG 10.5 17.87 8.3
3 AATAATATGAGTTAGATTGATA|TATCAATCTAACTCATATTATT 7.94 11.77 21.47
4 AGTAATTGTCGTTCTATCGATC|GATCGATAGAACGACAATTACT 5.08 7.47 13.04
5 ATATAAGCATAATATGGCTAAT|ATTAGCCATATTATGCTTATAT 5.01 7.55 15.15
6 CACACAATCACACAATCACACA|TGTGTGATTGTGTGATTGTGTG 4.72 8.57 2.32
7 ATTACTCTTATTATTATCAATC|GATTGATAATAATAAGAGTAAT 4.62 6.67 11.8
8 TCACACAATCACAATCACACAA|TTGTGTGATTGTGATTGTGTGA 3.76 7.01 1.54
9 ACAATTACTATACTTATTACTC|GAGTAATAAGTATAGTAATTGT 2.94 4.39 8.46
10 AGACAGAGACAGAGACAGAGAC|GTCTCTGTCTCTGTCTCTGTCT 2.17 5.66 1.03
11 CACAATCACGATCACACAATCA|TGATTGTGTGATCGTGATTGTG 1.43 2.25 0.5
12 CTGTCTCTGTCTGTCTCTGTCT|AGACAGAGACAGACAGAGACAG 1.34 3.77 0.68
13 CAGCGGATATGTGCGAATTAGA|TCTAATTCGCACATATCCGCTG 0.8 0.54 0.73
14 CTGAGCACAATTCAACACCACA|TGTGGTGTTGAATTGTGCTCAG 0.58 0.35 0.68
15 AACCTAACCTAACCTAACCTAA|TTAGGTTAGGTTAGGTTAGGTT 0.06 0.15 0.03
Location
/fs/szattic-asmg5/Bees/Megachile_rotundata/error_correction/large_libs/s_?_?_?kb.sequence.cor.all.txt /fs/szattic-asmg5/Bees/Megachile_rotundata/error_free/s_?_?_sequence.cor.txt /fs/szattic-asmg5/Bees/Megachile_rotundata/frg/ # frg files to assemble ginkgo:/scratch1/dpuiu/Megachile_rotundata/Data/error_free/noRepeats/ # repeat free FASTQ reads & FRG files
Assemblies
- CA Version: 6.1 (09/01/2010) /fs/szdevel/dpuiu/SourceForge/wgs-6.1/Linux-amd64/bin/runCA
- SOAP version 1.04: /nfshomes/dpuiu/szdevel/SOAPdenovo_Release1.04/
CA noOBT ; partial s_2_3kb, s_2_8kb, s_3
- Data : 3 libs : ~ 16X cvg
Gatekeeper
LibraryName numActiveFRG numDeletedFRG numMatedFRG readLength clearLength GLOBAL 72,995,448 0 70632632 8307194830 8278360381 LegacyUnmatedReads 0 0 0 0 0 s_2_3kb 9,166,343 0 8736228 942501164 914798596 s_2_8kb 210,266 0 199620 21669112 20742291 s_3 63,618,839 0 61696784 7343024554 7342819494
UID IID mateUID mateIID libUID libIID isDel isNonRandom Orient Length clrBeginLATEST clrEndLATEST 110000000001 1 120000000001 2 s_2_3kb 1 0 0 I 75 0 75 120000000001 2 110000000001 1 s_2_3kb 1 0 0 I 123 0 123 110000000003 3 120000000003 4 s_2_3kb 1 0 0 I 90 0 90 120000000003 4 110000000003 3 s_2_3kb 1 0 0 I 123 40 123 ... 110009166343 9166343 0 0 s_2_3kb 1 0 0 U 76 11 76 210009166344 9166344 220009166344 9166345 s_2_8kb 2 0 0 I 123 21 123 ... 210009376609 9376609 0 0 s_2_8kb 2 0 0 U 88 0 88 320009376610 9376610 0 0 s_3 3 0 0 U 72 0 72 ... 310072995448 72995448 0 0 s_3 3 0 0 U 68 0 68
BOG/ tigStore
- Number of tigs in the store
tigStore -g asm.gkpStore -t asm.tigStore 2 -D unitiglist | tail -1 | awk '{print $1}' # 36318422
- Single read tigs
tigStore -g asm.gkpStore -t asm.tigStore 2 -U -d layout | grep -c '^data.num_frags 1$' # 34985292 ts2lay | grep -B 9 -A 3 '^data.num_frags 1$'
Stats
. elem min q1 q2 q3 max mean n50 sum #repeats comments scf 20,827 122 3228 6374 13700 202495* 11508 20462* 239696810 SOAPdenovo: max=1102803 , N50=26876 ctg 37,494 65 2185 3998 7706 191323* 6380 10151* 239226293 206 SOAPdenovo: max=121554 , N50=3138 deg 1,136,469 64 123 143 184 5031 160 164 181954480 807132 utg 1,437,146 64 123 143 195 67048 308 870 443759899 readsTotal 72,995,448 readsInContigs 27,837,956 readsInDegenerates 9,627,122 singletons 34,881,692 (47%) readsWithOuttieMate 3,028,956(4.15%) ???
Placed reads . badLong badOuttie badSame bothDegen bothSurrogate diffScaffold good notMated oneChaff oneDegen oneSurrogate s_2_3kb 534 2,998,286 458 1614846 9892 21872 27308 267328 979980 760044 65268 s_2_8kb 4 26,864 10 38636 114 294 178 5044 35465 7848 1022 s_3 11072 3,806 1104 2369982 61236 53370 23058022 1208112 3967689 371538 87260
Chaff reads . bothChaff notMated oneChaff s_2_3kb 1,277,760 162,787 979,980 s_2_8kb 53,588 5,602 35,465 s_3 27,684,878 713,943 3,967,689
Issues
- reads are renamed : HWI-EAS385_0062:2:1:1036:15608#GCCAAT/1 => UID:110000000001 => IID:1
- reads < 64bp are deleted from the beginning : ID mapping ???
- lib s_2 orientation ??? Too many badOuttie's
Location
ginkgo:/scratch1/dpuiu/Megachile_rotundata/Assembly/wgs-noOBT-partial.1/
CA noOBT ; partial : s_2_3kb, s_2_8kb, s_3 ; no repeats
- Reads that contain at least one of the 15 most frequent 22mers are deleted from the input set
Gatekeeper
LibraryName numActiveFRG numDeletedFRG numMatedFRG readLength clearLength GLOBAL 33811292 0 32385550 3781368484 3772837304 LegacyUnmatedReads 0 0 0 0 0 s_2_3kb 5103461 0 4928188 518415011 510187775 s_2_8kb 53215 0 51436 5395700 5289866 s_3 28654616 0 27405926 3257557773 3257359663
Overlapper
- Dirty 3' ends for the s_2_* reads
totalOvl avgOvl s_2_3kb 5' 4955294 9 s_2_3kb 3' 4955294 7 s_2_8kb 5' 51050 10 s_2_8kb 3' 51050 7 s_3 5' 27721948 9 s_3 3' 27721948 9
Bog
cat 4-unitigger/asm.cga.0 | head
Global Arrival Rate: 0.125220
There were 1,983,199 unitigs generated.
Unitig Length
65407 - 67872: 4
50209 - 58608: 5
40073 - 49263: 27
30132 - 39913: 72
20030 - 29892: 319
10001 - 19992: 1979
9007 - 9999: 673
8000 - 8999: 934
7000 - 7999: 1332
6000 - 6998: 2048
5000 - 5999: 3103
4000 - 4999: 4898
3000 - 3999: 8120
2000 - 2999: 14634
1000 - 1999: 26621
900 - 999: 4116
800 - 899: 4457
700 - 799: 5042
600 - 699: 6146
500 - 599: 8107
400 - 499: 11901
300 - 399: 19373
200 - 299: 64394
100 - 199: 1173219
90 - 99: 161987
80 - 89: 189874
70 - 79: 132943
64 - 69: 82098
Stats
- Larger max scf & ctg !!! (compared with "CA noOBT partial" that assembled the repeats as well)
. elem min q1 q2 q3 max mean n50 sum scf 21041 65 3174 6334 13482 337719* 11376 20153* 239366537 ctg 37668 65 2181 3963 7687 191376* 6343 10083* 238928665 deg 380596 64 107 126 170 4688 163 160 62151395 utg 652051 64 115 133 225 67870 491 2469 320381694 readsTotal 33,811,292 readsInContigs 27,753,101 (82.08%) readsInDegenerates 4,004,853 (11.84%) singletons 1,276,811 (3.78%)
Placed reads . badLong badOuttie badSame bothDegen bothSurrogate diffScaffold good notMated oneChaff oneDegen oneSurrogate 1 582 2992742 410 773006 13486 20146 26870 159957 107838 753916 78412 2 1228 2 9486 84 124 62 1550 12940 7750 860 3 11354 3884 1074 2266364 90824 56066 23001316 1165421 346169 416116 156218
Chaff reads . bothChaff notMated oneChaff 1 52942 15316 107838 2 5932 229 12940 3 652176 83269 346169
~/bin/asm2mdi.pl < asm.asm s_2_3kb 16 87 ??? s_2_8kb 8000 800 s_3 337 27
Location
ginkgo:/scratch1/dpuiu/Megachile_rotundata/Assembly/wgs-noOBT-partial.1.noRepeats/
CA OBT ; partial : s_2_3kb, s_2_8kb, s_3 ; no repeats ; doDeduplication
Gatekeeper
LibraryName numActiveFRG numDeletedFRG numMatedFRG readLength clearLength GLOBAL 32121482 1689810 29520314 3627930463 3570353114 LegacyUnmatedReads 0 0 0 0 0 s_2_3kb 4600173 503288 4034304 473853527 454981975 s_2_8kb 47210 6005 41150 4851578 4628382 s_3 27474099 1180517 25444860 3149225358 3110742757
Stats
. elem min q1 q2 q3 max mean n50 sum scf 29488 70 2345 4369 8750 202300 7468.57 12354 220233106 ctg 60146 64 1480 2472 4394 77615 3645.31 5339 219251091 deg 294445 54 116 135 205 7625 200.63 218 59074721 utg 504418 52 121 150 320 63670 577.73 2333 291418460
CA noOBT ; partial s_3 , s_4 ; no repeats
- Reads that contain at least one of the 15 most frequent 22mers are deleted from the input set
Gatekeeper
LibraryName numActiveFRG numDeletedFRG numMatedFRG readLength clearLength GLOBAL 56839966 0 54032986 6425669702 6425397526 LegacyUnmatedReads 0 0 0 0 0 s_3 28654616 0 27405926 3257557773 3257359663 s_4 28185350 0 26627060 3168111929 3168037863
Stats
. elem min q1 q2 q3 max mean n50 sum scf 12908 148 4003 8811 22202 511831** 19416 42207** 250623581 ctg 23116 64 2888 5959 13088 255301** 10828 20109** 250302752 deg 274961 64 124 139 182 4652 172 164 47499725 utg 574873 64 123 132 202 128547 563 4478 324059992
CA noOBT
- Data : 7 libs : ~ 74X cvg
Gatekeeper
LibraryName numActiveFRG numDelFRG numMatedFRG readLength clearLength #repeats GLOBAL 326,236,387 0 315518526 37451489553 37418130441 LegacyUnmatedReads 0 0 0 0 0 s_2_3kb 9107424 0 9107424 942165284 910444046 # s_2_8kb 209336 0 209336 21814418 20787384 # s_3 63618839 0 61696784 7343024554 7342819494 # s_4 63544688 0 61255960 7291557748 7291478152 # s_5 63370860 0 61084368 7271218123 7271051639 # s_6 63780887 0 61685156 7359094156 7359012512 # s_7 62604353 0 60479498 7222615270 7222537214 #
Meryl
meryl -Dh -s 0-mercounts/asm-C-ms22-cm0 Found 30570218845 mers. Found 271464470 distinct mers. Found 11164787 unique mers. Largest mercount is 87984949; 1896 mers are too big for histogram. 1 11164787 0.0411 0.0004 2 9376915 0.0757 0.0010 3 3714582 0.0894 0.0013 ... 54 5344148 0.6573 0.1788 ...
fasta2tab.pl 0-mercounts/asm.nmers.ovl.fasta | sort -n -r | head -5 87,908,217 AATCATACAATCACAATCATAC 84,450,288 CAATCATACAATCACAATCATA ... 74,975,282 AATAATATGAGTTAGATTGATA
egrep -c 'AATCATACAATCACAATCATAC|GTATGATTGTGATTGTATGATT' *fastq *txt > egrep.count mulberry:/scratch2/dpuiu/Megachile_rotundata/Data/error_free/egrep.count
meryl -Dh -s 0-mercounts/asm-C-ms15-cm0 | head Found 32850820919 mers. Found 142500876 distinct mers. Found 2381895 unique mers. Largest mercount is 125816941; 2023 mers are too big for histogram. 1 2381895 0.0167 0.0001 2 2325770 0.0330 0.0002 3 708786 0.0380 0.0003 ... 54 1851586 0.4894 0.0671 ...
Overlap
- job count :
cat 1-overlapper/ovlopts.pl | grep ^\"h | wc -l 924
- Failures: 709 jobs failed; runCA 6.1 could not restart overlap properly !!!
cat 1-overlap/overlap*out | grep "^Could not" | sort -u Could not malloc memory (1305184948 bytes)
Bog
cat 4-unitigger/asm.cga.0
Global Arrival Rate: 0.443659
There were 158,805,551 unitigs generated.
Unitig Length
Global Arrival Rate: 0.443659
100071 - 168549: 21
90845 - 99102: 15
80566 - 88867: 17
70006 - 79485: 39
60191 - 69891: 51
50210 - 59643: 98
40106 - 49917: 191
30015 - 39986: 448
20006 - 29992: 1068
10000 - 19995: 4187
9001 - 9999: 942
8001 - 8998: 1202
7000 - 7999: 1489
6000 - 6999: 1927
5000 - 5999: 2379
4000 - 4999: 3266
3000 - 3999: 4580
2000 - 2999: 6979
1000 - 1999: 9654
900 - 999: 1176
800 - 899: 1346
700 - 799: 1658
600 - 699: 2405
500 - 599: 4742
400 - 499: 13047
300 - 399: 26578
200 - 299: 361389
100 - 199: 135260255
90 - 99: 7874207
80 - 89: 7147630
70 - 79: 5128367
63 - 69: 2427507
138,219,089 out of 158,805,551 contain one of the frequent kmers
CGW
- Monitor cgw
ps -C cgw PID PPID %MEM RSZ %CPU STIME TIME CMD 8563 8560 95.2 251872528 88.2 13:24 01:47:56 /fs/szdevel/dpuiu/SourceForge/wgs-6.1/Linux-amd64/bin/cgw ...
top -b -p 8563 -d 10 | grep dpuiu > cgw.resource_usage.log
- Failure 1:
tail 7-0-CGW/cgw.out ... Processed 158,288,858 unitigs with 326,296,236 fragments #Bumble bee : Processed 61,930,044 unitigs with 301,738,113 fragments * Loaded dist s_2_3kb,1 (3000 +/- 300) * Loaded dist s_2_8kb,2 (8000 +/- 800) * Loaded dist s_3,3 (475 +/- 47.5) ... * Splitting chimeric input unitigs LIB 1 mu = 15.318100 sigma = 89.035478 LIB 2 mu = 8000.000000 sigma = 800.000000 LIB 3 mu = 337.817628 sigma = 26.699549 ... minLength = 460 minSplit = -429 Splitting unitig 47689 into as many as 3 unitigs at intervals: 22905,22906 .. Splitting unitig 158234882 into as many as 3 unitigs at intervals: 124,136 * BuildGraphEdgesDirectly
Fix (partial): add "-I" flag to cgw in runCA cat 7-0-CGW/cgw.out ... *** BuildGraphEdgesDirectly Operated on 171664374 fragments
- Failure 2:
tail 7-0-CGW/cgw.out **** Calling CheckEdgesAgainstOverlapper **** **** Survived CheckEdgesAgainstOverlapper with 0 failures**** * Allocating Contig Graph with 158289029 nodes and 14055921 edges Could not calloc memory (25326244640 * 1 bytes = 25326244640) cgw: AS_UTL_alloc.C:55: void* safe_calloc(size_t, size_t): Assertion `p != __null' failed.
Location
mulberry:/scratch2/dpuiu/Megachile_rotundata/Assembly/wgs-noOBT
SOAPdenovo (Tanja)
cat *.ContigIndex | grep -v ^E | grep -v ^i | count.pl -i 1 | getSummary.pl -j 1 -t "contigs" cat *.ContigIndex | grep -v ^E | grep -v ^i | count.pl -i 1 | getSummary.pl -j 1 -min 100 -t "contigs(>100bp)" grep "^>" *.scaf | getSummary.pl -i 2 -t scaf
- Stats
. elem min q1 q2 q3 max mean n50 sum contigs 9742349 31 32 33 37 114832 60 44 585430821 contigs(>100bp) 177327 100 131 261 1398 114832 1333 3897 236496823 # N50 for Bee was 7K scaf 7863 102 903 3272 17692 2338728 37825 240706 297423517 # N50 for Bee was 1.17M
- Location
/fs/szattic-asmg5/Bees/Megachile_rotundata/Assembly/assembly5kbForAll
SOAPdenovo (Daniela)
Stats
cat asm.K31.contig | grep "^>" | awk '{print $3}' | uniq -c | awk '{print $2,$1}' > asm.K31.contigLen.count
. elem min q1 q2 q3 max mean n50 sum scaff 25,119 351 1896 4444 10914 1,102,803 11041 26876 277,338,897 contigs(all) 6,917,796 31 32 34 40 121,554 70 73 487,401,812 contigs(>100bp) 210,666 100 124 222 1174 121,554* 1108 3138* 233,563,401
reads 340,437,486 readsOnContigs 171,212,613
Alignments
- Align the 3kb & 8kb libs to the scaffolds
soap2-index asm.K31.scafSeq mkdir soap2-index mv asm.K31.scafSeq.index.* soap2-index/ soap2 -D ... -a s_2_1_3kb_sequence.txt -b s_2_2_3kb_sequence.txt -l 32 -p 16 -v 2 -m 2000 -x 4000 -o s_2_3kb.mated.soap2 -2 s_2_3kb.single.soap2 -R soap2 -D ... -a s_2_1_5kb_sequence.txt -b s_2_2_5kb_sequence.txt -l 32 -p 16 -v 2 -m 4000 -x 6000 -o s_2_5kb.mated.soap2 -2 s_2_5kb.single.soap2 -R soap2 -D .. -a s_2_1_8kb_sequence.txt -b s_2_2_8kb_sequence.txt -l 32 -p 16 -v 2 -m 6000 -x 10000 -o s_2_8kb.mated.soap2 -2 s_2_8kb.single.soap2 -R soap2 -D .. -a s_3_1_sequence.txt -b s_3_2_sequence.txt -l 32 -p 16 -v 2 -m 200 -x 400 -o s_3.mated.soap2 -2 s_3.single.soap2 ...
mates mated single single.diffScaff single.sameScaff s_2_3kb 21,563,283 1,545,114 8,449,321 341,974 1,203,570 s_2_5kb 36,218,589 5,639,332 44,533,553 4,784,348 30,621,038 s_2_8kb 198,377 1,068 32,280 1,168 2,842 s_2_3kb.filter 4,823,235 3,730 3,618,426 38,562 3,017,172 s_2_8kb.filter 111,267 20 33,819 372 27,300 s_3 35,548,153 15,521,020 4,589,276 37,564 651,924 s_2_1kb 1,466,112 8,014,900 84,970 191846 lib mates mated% single% single.diffScaff% single.sameScaff% s_2_3kb 21,563,283 3.58 19.59 0.79 2.79 s_2_5kb 36,218,589 7.785 61.475 6.6 42.27 s_2_8kb 198,377 0.265 8.135 0.29 0.715 s_2_3kb.filter 4,823,235 0.035 37.51 0.395 31.275 s_2_8kb.filter 111,267 0.005 15.195 0.165 12.265 s_3 35548153 21.83 6.455 0.05 0.915
Location
mulberry:/scratch2/dpuiu/Megachile_rotundata/Assembly/SOAPdenovo-redo
SOAPdenovo ; partial : s_[34567] ; no repeats
- Similar results to SOAPdenovo : wrong inserts & repeats don't affect much
Stats
. elem min q1 q2 q3 max mean n50 sum
scf 24602 333 1724 4380 11049 1,103,462 11135 27887 273963709 contigs(all) 2515516 31 33 36 52 148,198 131 1880 330512911 contigs(100bp+) 184395 100 127 235 1316 148,198 1263 3730 232932308