Trace formatting
Articles
- The impact of next-generation sequencing technology on genetics Elaine Mardis Trends in Genetics 2008
- As Users Demand Paired-End Sequencing, 454, Illumina, and ABI Work On New Kits
- Sequence Format Descriptions(EMBL)
- ismb2007Poster.pdf
- Smith_Rennes_2007.pdf
- GenomeAnalyzer_SpecSheet.pdf
- Assembly and Alignment Algorithms for Next-Gen Sequence Data
- Sequencing of natural strains of Arabidopsis thaliana with short reads (Illumina)
- Accuracy and quality of massively parallel DNA pyrosequencing
- Advanced sequencing technologies and their wider impact in microbiology
- Emerging technologies in DNA sequencing
- Lander Waterman
- Whole-genome re-sequencing
- Whole-Genome Sequencing and Assembly with High-Throughput, Short-Read Technologies
- seqanswers
- politigenomics table
- Valex & SRA
- BIOINFORMATICS FOR NEXT GENERATION SEQUENCING
- The Short Read Headache
- The impact of next-generation sequencing technology on genetics
- Next-generation DNA sequencing methods.
- Short-Read Sequencing Technologies for Transcriptional Analyses.
- Next Generation Sequencing - Bioinformatics Journal - Virtual Edition
Cleaning
- run dust on Ecoli & UniVector to mask low complexity seqs
- screen against Ecoli & UniVector database
Technologies
Latest Technology Summary
Technology 454 Illumina Solid
seq-by-synthesis seq-by-synthesis ABI Company 454(Roche) Illumina Location Brandford,CT SanDiego,CA Latest GS FLX, Titanium reagents Genome Analyzer II SOLID 3
Throughput 500M/run 1G/run 20G/run RunTime 10hr 3days ReadLen 500bp 36 35 InsertLen 3K 100-200bp 600-10K
Accuracy 99.94%
Q20(99%accuracy) 400bp 34bp
Cost $3K/run 60K/3G
$400/4M bacterial genome(25-30X)
Problems homopolimers
Sanger
454
- Pyrosequencing
- 454_Life_Sciences wikipedia
- In-sequenece article
- 1_paired_end.pdf
- [1]
- [2]
Anomalies:
* homopolymer lengths can be shorter than real * substitutions less likely than in traditional methodssingle base insertions * carry forward events usually near but not adjacent to homopolymers
GS20
* 1.6M total wells * 450K detactable wells * 200K usable wells
Accuracy: * published per-base accuracy of a Roche GS20 is only 96%. * Mitch Sogin paper * 99.5% accuracy rate in unassembled sequences * identified several factors that can be used to remove a small percentage of low-quality reads, improving the accuracy to 99.75% or better => better quality than Sanger sequencing * The error rate, defined as the number of errors (miscalled bases plus inserted and deleted bases) divided by the total number of expected bases, was 0.49% * 36% insertions, 27% delitions, 21% N's, 16% substitutions * A to G and T to C, were more frequent than other mismatches * reverse transitions, G to A and C to T, were not that frequent * Nearly 70% of the homopolymer extensions were A/T * errors were evenly distributed along the length of the reference sequences, they were not evenly distributed
among reads: 82% had no errors, 93% had no more than a single error, and 96% had no more than 2 errors.
* A small number of reads, fewer than 2%, contained a disproportionate number of errors that account for nearly 50% of the miscalls for the entire dataset * Avg quality is 25; in homopolymers can drop as low as 5 * Reads much longer than avg length had more errors * strong correlation between the presence of ambiguous base calls and other errors in a read * The presence of even a single ambiguous base in a read correlates strongly with the presence of other errors * Primer errors also correlated with errors
GS FLX
GS FLX with Titanium reagents
* up to 500M/run * reads up to 500bp
Get info from .sff files:
$ sffinfo -h
Usage: sffinfo [options...] [- | sfffile] [accno...]
Options:
-a or -accno Output just the accessions
-s or -seq Output just the sequences
-q or -qual Output just the quality scores
-f or -flow Output just the flowgrams
-t or -tab Output the seq/qual/flow as tab-delimited lines
-n or -notrim Output the untrimmed sequence or quality scores
-m or -mft Output the manifest text
un-paired reads
paired ends
Features:
* approximately 84-nucleotide DNA fragments * have a ~ 44-mer linker sequence in the middle * flanked by a ~ 20-mer sequence on each side. * The two flanking 20-mers are segments of DNA that were originally located approximately 2.5 (3?) kb apart in the genome of interest. * The ordering and orienting of contigs generates scaffolds which provide a high-quality draft sequence of the genome.
Linker(palindrome) : GTTGGAACCGAAAGGGTTTGAATTCAAACCCTTTCGGTTCCAAC Check for linker : sffinfo -s *.sff | ~/bin/fasta2tab.pl | grep GTTGGAACCGAAAGGGTTTGAATTCAAACCCTTTCGGTTCCAAC 12345678901234567890123456789012345678901234 GTTGGAACCGAAAGGGTTTGAATTCAAACCCTTTCGGTTCCAAC
GTTGGAACCGA AAGGGTTTGAA TTCAAACCCTT TCGGTTCCAAC
Anomalies:
* the linker can appear (tandem,completely/partially) more than once * some reads end up in linker (partial) * some reads don't contain the linker at all * some reads are cloning vector
NCBI bacterial data sets:
Accession Center Instrument SRR001351,2,3 JCVI 454 GS FLX 454 Paired End Sequencing Porphyromonas gingivalis W83 SRR001355 JCVI 454 GS FLX 454 Paired End Sequencing Escherichia coli str. K-12 MG1655 SRR004895,9 BI 454 GS FLX 45X on 454 (15X fragment and 30X paired) Brucella pinnipedialis SRR004900,1 BI 454 GS FLX 45X on 454 (15X fragment and 30X paired) Brucella suis bv. 3 SRR005309,10,11 BI 454 GS FLX 45X on 454 (15X fragment and 30X paired) Brucella ceti SRR005481,2 BI 454 GS FLX 45X on 454 (15X fragment and 30X paired) Brucella ceti BROAD:SEQUENCING_SAMPLE:24613.2 SRR005486,7,8 BI 454 GS FLX 45X on 454 (15X fragment and 30X paired) Brucella pinnipedialis BROAD:SEQUENCING_SAMPLE:246 SRR006465 GSC 454 GS FLX 454 Paired-End Library. Acinetobacter sp. ADP1
File location:
/fs/szdata/454p/ /fs/szasmg2/Bacteria/Pseudomonas_syringae/Data/DC3000.format.454Reads.fna : Pseudomonas_syringae
Illumina
- Sequencing by Synthesis
- Platforms:
* Genome Analyzer (GA) * Genome Analyzer II : faster, higher tput * Future: 10GB/run 50bp reads * Future: 20GB/run 100bp reads
Data sets:
Strep suis Solexa data set for download at Sanger Staphylococcus aureus strain MW2 (edena paper) NCBI Solexa example data set Pseudomonas aeruginosa Pseudomonas syringae NCBI SRA
Applications:
* Gene Expression * ChIPSeq (hight throughput) * Re-sequencing * mRNA sequencing
Software:
Staden & Io_lib * IO_LIB package /fs/sz-user-supported/common/packages/io_lib-1.11-x86_64/bin/ * STADEN package /fs/sz-user-supported/common/packages/staden-src-1-7-0/distrib/unix-rel-1-7-0/linux-bin MAQ Sanger assembler FASTQ sequence format
Illumina 1G :
* ~40 Million DNA sequencing reactions * about 36 hours for a run * each sequence is up to 36 bases long * insert len=~200bp
Illumina Genome Analyzer II:
* up to 51 bp * mate-pairs: opposite directions, slight overlap (insert size is less than 200bp "advertised") * on the SRA mate-pairs are joined; when downloaded only one read is shown. What about the mate pair?
SRA: set of 4 files
*_seq.txt : lane,run, well(x,y) sequence *_prb.txt : max quality from each group of 4 values is taken as quality *_sig2.txt : lane,run, well(x,y); max signal from each group of 4 values corresponds to max quality *_qhg.txt : lane,run, well(x,y); some encoded info?
# *_seq.txt 5 1 1269 1795 AAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA # _prb.txt 40 -40 -40 -40 40 -40 -40 -40 ... # _sig2.txt <== 5 1 1269 1795 2594.0 2367.0 -10.0 -96.0 ...
Qualities:
Range : -5..40 Avg : ~25, depending on the data set
Fastq format
Example:
1 lane of Solexa reads: 10,959 READS; all are 36 bp $ /fs/sz-user-supported/common/packages/io_lib-x86_64/bin/solexa2srf s_8_0100_seq.txt ; mv traces.srf s_8_0100.srf $ /fs/sz-user-supported/common/packages/io_lib-x86_64/bin/srf2fastq s_8_0100.srf > s_8_0100.fastq @s_8_100_293_551 CCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCACC + IIIIIIIIIIIIIIIIIIIIIIIIIIIIIIIIIIII @s_8_100_35_698 TATATGATTGACAATATAAAAATATGAGTATAAAAT + IIIIIIIIIIIIIIIIIIIIIIIIIIIIIIII4/:I @s_8_100_880_947 TTATTATCTTTATTGACGTACCTCTAGAAGACCCAA + IIIIIIIIIIIIIIIIIIIIIIIIIIIIIIIII;>1 ...
Edge effect: N's have quality -14
$ cat s_8_0100_seq.txt | sort -nk3 -nk4 8 100 0 37 ......AT.AT...TAATCAATA..GA.GAAG.... ... 8 100 1003 959 AGTC.......T.C.........GT.........AA
$ more traces.qual ... >s_8_100_0_37 -14 -14 -14 -14 -14 -14 25 13 -14 25 25 -14 -14 -14 25 25 25 25 22 25 25 25 25 -14 -14 25 25 -14 25 -11 25 14 -14 -14 -14 -14 ... >s_8_100_1003_959 25 25 25 25 -14 -14 -14 -14 -14 -14 -14 25 -14 25 -14 -14 -14 -14 -14 -14 -14 -14 -14 25 -10 -14 -14 -14 -14 -14 -14 -14 -14 -14 8 25 ...
# bioperl script to convrt seq formats
$ seqconvert.PLS --from fastq --to fasta < s_8_0100.fastq
# get fastq qualities
$ more *fastq | grep -A 1 "^+" | grep -v ^+ | grep -v -- ^-- | perl -ane '@F=split //,$F[0]; foreach (@F) { $n=ord($_)-33; print $n," ";} print "\n";'
# convert Solexa format (maq fq_all2std.pl script) $ fq_all2std.pl seqprb2std s_5_0001_seq.txt s_5_0001_prb.txt > s_5_001.fastq $ fq_all2std.pl fq2fa s_5_001.fastq > s_5_001.seq
# convert fastq to seq & qual fastq2seqqual.pl test.fastq fastq2seqqual.pl -solexa test.fastq => test.seq , test.qual
SOLiD
- ABI SOLiD
- article
- Tools & Data Sets
- color space (0123) => base space (ACGT)
- .csfsta file : in color space; start with a known base (usually T)
Example:
Ecoli_F3.csfasta >600_50_31_F3 T2222002113300322132112231 >600_50_63_F3 T2330133212130133221033110
Ecoli_F3_QV.qual >600_50_31_F3 15 20 14 11 25 20 17 21 16 9 15 12 21 8 2 10 15 5 3 5 10 6 2 4 4 >600_50_63_F3 5 13 9 23 5 9 8 4 6 4 5 4 7 6 10 7 7 5 13 11 5 8 2 7 6
Ecoli_R3.csfasta >600_51_85_R3 G0331332123123330101312331 >600_51_178_R3 G1111033111110111101111111
Ecoli_R3_QV.qual >600_51_85_R3 10 13 16 15 11 17 6 13 10 9 15 8 13 10 12 11 8 5 3 6 12 8 11 6 14 >600_51_178_R3 2 2 2 5 8 2 2 2 2 3 2 3 2 3 7 4 3 4 5 4 2 2 5 6 17
- low error rate (higher accuracy than Illumina)
- 2008: 4G run, read_len=35bp; insert=3Kbp (old)
- 2009: 9G run, read_len=50bp;
- SOLiD™ 3 System generates (Oct 1 2008)
- over 20 gigabases
- mate-paired libraries with insert sizes ranging from 600 bp up to 10 kbp
- human genome for less than $60,000.
- uniform bases quality
- accuracy greater than 99.94%
- because of double base interogation & high cvg, qualities can be "discarded"
Alignment matrix
A C G T A 0 1 2 3 C 1 0 3 2 G 2 3 0 1 T 3 2 1 0
Examples:
AA is encoded as 0 CG is encoded as 3 AACG is encoded as 0 1 3
Features of Color space:
* Color space data are self-complementary
Example:
Base A G C T C G T C G T G C A G
Color space 2 3 2 2 3 1 2 3 1 1 3 1 2
Complemented
Base T C G A G C A G C A C G T C
Color space 2 3 2 2 3 1 2 3 1 1 3 1 2
* Two-Base Encoding and Error Recognition 1 change: measuring error multiple changes starting at a certain point: SNP
Example:
Reference 2 3 2 2 3 1 2 3 1 1 3 1 2
Observed 2 3 2 2 0 1 2 3 1 1 3 1 2
Data Sets:
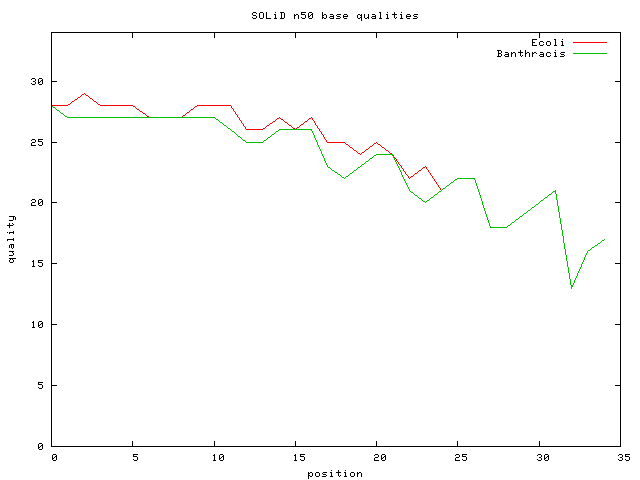
- E.Coli DH10B Mate-Pair Data Set ~ 56M 25bp paired reads =~ 1G bp ; 56M*25/4.6M => 300X cvg
- E.Coli DH10B Mate-Pair Data SubSet ~ 1.9M 25bp paired reads ; 10.17X cvg; avg inset is 2Kbp
- Bacillus anthracis 27M 35 bp unmated reads => 187X cvg
Helicos
- Web site
- @Broad
- 100Mbp/hour, 33bp reads, 5% error rate
- UMD instrument
Pacific Biosciences
- http://www.pacificbiosciences.com/index.php
- http://www.bio-itworld.com/BioIT_Content.aspx?id=71746
- Science paper
Visigen
Polonator
RainDance Technologies
Download
Genbank
- Bioperl scripts:
bp_fetch.pl net::genbank:NC_005810.1 > NC_005810.1 bp_fetch.pl net::genbank:NC_005810 > NC_005810 bp_fetch.pl net::genbank:45439865 > 45439865
TA
SRA
Format
TA
- Sanger:
tarchive2amos -o Ba Ba.seq # TA FTP tarchive2amos -o Ba -tracedir traces/ # TA querytrace_db tarchive2amos -o Ba -assembly assembly/ASSEMBLY.xml -tracedir traces/ # AA
SRA
- Solexa mated:
seq2amos.pl -n solexap -m 100 -s 20 -fs solexa_f.fasta -rs solexa_f.fasta > solexap.afg seq2amos.pl -n solexap -m 100 -s 20 -fs solexa_f.fasta -fq solexa_f.qual -rs solexa_r.fasta -rq solexa_r.qual > solexap.afg seq2amos.pl -n solexap -m 100 -s 20 -fs solexa_f*.fasta -rs solexa_r*.fasta > solexap.afg
- Solexa unmated:
seq2amos.pl -n solexa -fs solexa.fasta > solexa.afg
- 454 mated:
seq2amos.pl -n 454p -m 3000 -s 300 -fs 454.seq > 454p.afg
- 454 unmated:
seq2amos.pl -n 454 -fs 454.seq > 454.afg
Convestion
- Readseq
~/bin//readseq.sh -f Fasta -o prefix.fasta prefix.embl
- Bioperl
bp_sreformat.pl -i prefix.embl -o prefix.fasta -if EMBL -of Fasta
- AMOS:
amos2frg [-i infile] [-o outfile] amos2sq [-i] infile [-o outprefix] => outprefix.seq outprefix.qual
Read Mapping Software
BFAST
- need to e-mail to author to get the code
BLAT
- BLAT—The BLAST-Like Alignment Tool, Genome Research 2002
- FAQ
- Can align any type of reads
- Can do nt:aa translation
- Command: blat
blat -noHead -t=dna -q=dna -tileSize=10 -stepSize=3 Pa.1con Pa.seq Pa.blat
MAQ *
- Maq Sourceforge
- Maq Poster from Sanger
- Illumina-Solexa/AB-SOLiD , not 454 or capillary reads
- Uses FASTQ format
- Command: maq map ...
- does ungapped alignment on unpaired reads
SOLEXA maq.pl easyrun -d . ref.1con reads.fastq
SOLID solid2fastq.pl reads_ shortname maq fastq2bfq shortname.fastq shortname.bfq maq fasta2csfa ref.fasta > ref.csfa maq fasta2bfa ref.csfa ref.csbfa maq fasta2bfa ref.fasta ref.bfa maq map -c aln.cs.map ref.csbfa shortname.bfq 2> aln.log maq csmap2nt aln.nt.map ref.bfa aln.cs.map maq assemble cns.cns ref.bfa aln.nt.map 2> cns.log
RMAP
- RMAP : designed for Illumina-Solexa
- Command: rmap
rmap -m 3 -w 33 -c Pa.1con Pa.seq -o Pa.rmap
SHRiMP
- Web site
- Commands: rmapper-cs , rmapper-ls, ...
SeqMap
- SeqMap developed at Stanford
- allows up to five mixed substitutions and inserted/deleted nucleotides in the mapping
- allows sequences to contain N’s, and to have unequal lengths
./seqmap Usage: seqmap <number of mismatches> <probe FASTA file name> <transcript FASTA file name> <output file name> [options] Parameters: <number of mismatches> maximum edit distance allowed <probe FASTA file name> probe/tag/read sequences <transcript FASTA file name> reference sequences <output file name> name of the output file ...
SHORE
SOAP *
- Web site (China)
- Formatofoutput
- SOAP: short oligonucleotide alignment program, Bioinformatics Jan 2008
- Commands: soap, soap.contig, soap_dealign, soap.huge, soap.short
- can use qualities, do read trimming, use pair ends, RNA alignments
soap -v 5 -d Pa.1con -a Pa.seq -o Pa.soap
SOCS
- Web site
- ABI color space
socs socs.pref more socs.pref Req.fa Seq_F3.csfasta Seq_F3_QV.qual out_prefix 2 1000 2 false true 0
SOLiD
SSAHA
- Web site(Sanger)
- Focused on exact, nearly exact matches
- Does not find all the exact matches???
- Example: Solexa 33bp ~30% of reads are not found
ZOOM
Genome Resequencing
- The complete genome of an individual by massively parallel DNA sequencing (J.Watson's genome) Nature April 2008
- J.Watson's genome (supplementary info)
Links
Data
Solexa
- Strep suis Solexa at Sanger 36bp, ~49X coverage
- Staphylococcus aureus strain MW2 (edena paper) 35bp, ~47X coverage
- Pseudomonas aeruginosa: 33bp, ~43X coverage
- Pseudomonas syringae: 32bp, ~31X coverage
- 1000 Genomes (June 14th 2008): 47bp
Accession #Runs Instrument Center Study [Individual] SRA000303 41 Solexa 1G Genome Analyzer BI 1000Genomes Project Pilot 2 NA12878 SRA000304 49 Solexa 1G Genome Analyzer BI 1000Genomes Project Pilot 2 NA12891 SRA000305 56 Solexa 1G Genome Analyzer BI 1000Genomes Project Pilot 2 NA12892 SRA000307 1 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 1 NA10851 SRA000308 2 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 1 NA11993 SRA000309 3 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 1 NA11995 SRA000310 1 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 1 NA12006 SRA000311 1 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 1 NA12044 SRA000312 2 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 1 NA12156 SRA000313 1 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 1 NA12414 SRA000314 1 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 1 NA12776 SRA000315 1 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 1 NA12828 SRA000316 12 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 2 NA12878 SRA000317 8 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 2 NA12891 SRA000318 14 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 2 NA12892 SRA000319 1 Solexa 1G Genome Analyzer SC 1000Genomes Project Pilot 1 NA12004
June 14th 2008: Sept 19th 2008
SRA001100 23 Illumina Genome Analyzer BGI 1000Genomes Project Pilot 2 NA19240 ... SRA002029 1 Illumina Genome Analyzer II WUGSC 1000Genomes Project Pilot 2 NA19239
/fs/szdata/Solexa/1000genomes
- Example SRR001113.seq :
7,058,926 47 bp sequences 2,402,398 contain at least 1 '.'
454
- 1000 Genomes
June 14th 2008
Accession #Runs Instrument Center Study [Individual] SRA000302 121 454 GS FLX BCM 1000Genomes Project Pilot 2 NA12878 SRA001032 2 454 GS FLX BCM 1000Genomes Project Pilot 2 NA12878 SRA001036 1 454 GS FLX BCM 1000Genomes Project Pilot 1 NA12812 SRA001094 1 454 GS FLX BCM 1000Genomes Project Pilot 2 NA12878
June 14th 2008: Sept 19th 2008
SRA001037 2 454 GS FLX BCM 1000Genomes Project Pilot 1 NA12812 ... SRA001819 1 454 GS FLX BCM 1000Genomes Project Pilot 2 NA12878
- Cryptosporidium_muris_RN66: SRA001029 (not paired)
- EcoliK12: SRR001355 (paired)
- Porphyromonas_gingivalis_W83: E8YURXS01 (paired)
Refseq
- /fs/szdata/genomes/human_ncbi_build36/ NCBI build36.1 May 2006 (Current build is 36.3 March 2008)
- /fs/szdata/genomes/human_celera_2001_Orig/
Alignments
Whole genomes alignments
- BLAT
- BLASTZ, Post-processing long pairwise alignments article. decom program
- LAGAN
- MUMMER
- AVID